Epigenomics Sequencing
What Are Epigenetic Modifications
Epigenetic modifications are reversible chemical changes that regulate gene expression without altering the underlying DNA sequence. These modifications can be transmitted during cell division and play a crucial role in controlling gene activity. The most extensively studied epigenetic mechanisms include DNA methylation and chromatin modifications, both of which are essential for gene regulation and cellular function.
Epigenetic regulation participates in many biological processes such as cell differentiation, organism development, and tumor formation. DNA methylation typically occurs at the C5 position of cytosine within CpG dinucleotides in vertebrates, while non-CpG methylation (CHG and CHH) is commonly observed in plants and mammalian embryonic stem cells. These methylation patterns are established and maintained by DNA methyltransferase enzymes, mainly DNMT1, DNMT3A, and DNMT3B.
Main Components of Epigenomics
DNA Methylation Sequencing Technologies
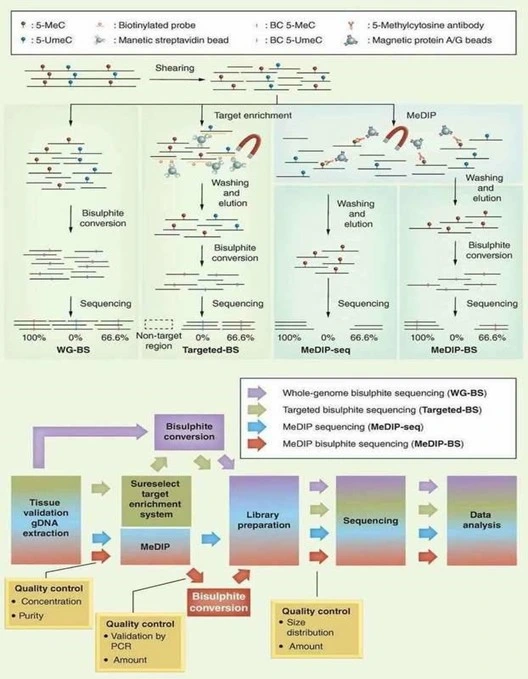
DNA methylation patterns can be lost during conventional molecular biology procedures such as PCR amplification or molecular cloning, because these processes do not preserve the activity of DNA methyltransferases responsible for maintaining methylation marks. To overcome this limitation, several specialized techniques have been developed to preserve methylation information and convert it into quantifiable genomic data.
Among the most widely used methods are methylated DNA immunoprecipitation (MeDIP), bisulfite sequencing (BS-seq), and reduced representation bisulfite sequencing (RRBS). These approaches enable the detection and measurement of DNA methylation across the genome. When combined with high-throughput sequencing (NGS) technologies, they provide comprehensive and reliable genome-wide methylation profiles.
Each technique has specific advantages and limitations. Bisulfite sequencing (BS-seq) and RRBS allow base-resolution analysis of the DNA methylome, enabling precise identification of methylated cytosines. In contrast, MeDIP-seq detects relative enrichment of methylated regions rather than exact methylation sites.
In addition to DNA methylation analysis, chromatin immunoprecipitation (ChIP) is widely used to study histone methylation patterns associated with specific gene promoters and to compare epigenetic states between normal and diseased tissues. The integration of ChIP with next-generation sequencing, known as ChIP-seq, enables genome-wide identification of DNA-binding protein targets and provides insights into protein–DNA interactions that regulate gene expression and cellular processes. This technology allows comprehensive chromatin analysis with high sensitivity and minimal sequencing bias.
Detection of RNA Methylation
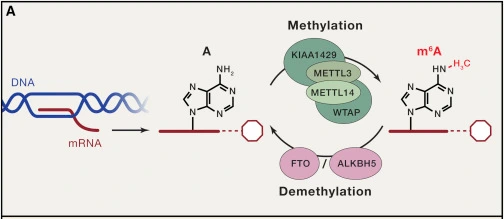
RNA methylation, similar to DNA methylation, involves the enzymatic addition of a methyl group (CH₃) to specific nucleotides within RNA molecules. This reaction is catalyzed by methyltransferase enzymes. More than 100 different chemical RNA modifications have been identified in cells, including m6A, m5C, m1A, and m7G methylation. Among these, N6-methyladenosine (m6A) is the most abundant and well-characterized modification in eukaryotic mRNA. It corresponds to the methylation of the nitrogen atom at the 6th position of adenine, representing nearly 80% of RNA methylation events and playing an essential role in post-transcriptional gene regulation.
Several experimental approaches have been developed to detect RNA methylation:
1. MeRIP-seq :
This technique relies on antibody-based enrichment of methylated RNA fragments. Antibodies specific to m6A bind to methylated RNA regions, which are then isolated and sequenced using high-throughput sequencing technologies. Although MeRIP-seq enables genome-wide identification of methylated regions, it generally detects methylation peaks rather than single-nucleotide resolution.
2. miCLIP (Methylation Individual-Nucleotide Resolution Cross-Linking and Immunoprecipitation):
miCLIP improves resolution by cross-linking antibodies to methylated RNA using ultraviolet light. During reverse transcription, cross-linked sites generate mutations or truncations in cDNA, which mark the exact location of methylated nucleotides. This method allows single-base resolution mapping of RNA methylation, but its application can be limited due to high cost and technical complexity, including the use of isotopic labeling.
Nanopore sequencing represents a third-generation sequencing technology capable of detecting RNA modifications directly. As RNA molecules pass through a nanopore channel, each nucleotide generates a characteristic electrical signal. Modified bases produce distinct signal patterns, enabling real-time identification of both nucleotide sequences and chemical modifications, including RNA methylation, at single-nucleotide resolution without the need for antibodies or chemical conversion.
Advantages of Epigenomics Sequencing
1. Comprehensive Genome-Wide Insight
Epigenomic sequencing provides a global view of epigenetic modifications across the entire genome, including DNA methylation, histone modifications, and regulatory non-coding RNAs. This integrated perspective helps researchers understand complex gene-regulatory networks and cellular responses.
2. High Sensitivity and Specificity
Modern epigenomic techniques such as ChIP-seq, Bisulfite sequencing, and ATAC-seq offer high sensitivity and accuracy. These methods allow precise detection and quantification of subtle epigenetic changes, which is essential for studying gene regulation and disease mechanisms.
3. High-Throughput Capability
Epigenomic sequencing technologies enable the simultaneous analysis of thousands of genomic regions. This high-throughput capacity improves research efficiency and supports large-scale genomic and epigenetic studies.
4. Temporal and Cellular Resolution
Advanced epigenomic tools can analyze epigenetic modifications across different developmental stages and cell types. This allows scientists to investigate the dynamic nature of epigenetic regulation in development, differentiation, and disease progression.
Applications of Epigenomics Sequencing
Epigenomics sequencing is widely used across multiple scientific disciplines to understand gene regulation and cellular function. In cancer research, it helps identify epigenetic alterations in tumor cells, enabling the discovery of biomarkers and potential therapeutic targets that support personalized medicine. In developmental biology, epigenomic studies reveal how epigenetic mechanisms regulate embryonic development, cell differentiation, and organ formation. In neuroscience, epigenomic analysis provides insights into the role of epigenetic modifications in neurodevelopment, synaptic plasticity, and neurodegenerative diseases, helping to explain the molecular basis of neurological disorders. Epigenomics is also valuable in environmental science, where it is used to investigate how external factors such as diet, pollutants, and lifestyle influence gene expression and epigenetic regulation. Additionally, in plant science, epigenomic approaches help uncover the molecular mechanisms controlling plant growth, development, and responses to environmental stress, contributing to crop improvement and agricultural productivity.