Gene Regulation By miRNA
Introduction
MicroRNAs (miRNAs) are short, single-stranded, non-coding RNA molecules, roughly 22 nucleotides long, that play key roles in regulating gene expression. First discovered in the early 1990s, miRNAs have since been shown to influence many biological processes, including development, cell proliferation, differentiation, apoptosis, and metabolism. They function by binding to complementary sequences in the 3′ untranslated regions (3′UTRs) of target messenger RNAs (mRNAs), resulting in either translation suppression or mRNA degradation.
Research on miRNAs has revealed fundamental insights into biology and offers promising avenues for new therapeutic strategies across multiple diseases. This review highlights recent advances in miRNA biology, covering their biogenesis, mechanisms of action, and roles in development, cell growth, apoptosis, and metabolism. Additionally, it explores emerging technologies for potential miRNA-based therapies.
miRNA Biogenesis
miRNAs are initially transcribed in the nucleus by RNA polymerase II as long primary transcripts (pri-miRNAs) containing hairpin structures. These pri-miRNAs are processed by the microprocessor complex, which includes Drosha and DGCR8, into precursor miRNAs (pre-miRNAs) of ~70 nucleotides with stem-loop structures. Pre-miRNAs are then exported to the cytoplasm via the exportin-5/Ran-GTP complex. In the cytoplasm, the RNase III enzyme Dicer cleaves pre-miRNAs to produce mature miRNAs, though some miRNAs, like miR-451, are processed independently of Dicer by Argonaute 2.
Mature miRNAs (21–23 nucleotides) are incorporated into the RNA-induced silencing complex (RISC), where they guide the complex to complementary sequences in the 3′ untranslated regions (3′UTRs) of target mRNAs, leading to mRNA degradation or translational repression.
miRNA biogenesis is tightly regulated at multiple levels. Transcription is controlled by transcription factors and epigenetic modifications, while Drosha and Dicer processing is influenced by protein interactions and RNA-binding proteins. miRNA stability and activity can also be modulated by epigenetic modifications and competing endogenous RNAs (ceRNAs), which sequester miRNAs and limit their interaction with target mRNAs.
miRNAs and Biological Processes
miRNAs regulate gene expression by binding to complementary sequences in the 3′ untranslated regions (3′UTRs) of target mRNAs, leading to either mRNA degradation or translational repression, depending on the degree of complementarity. They can also interact with other regions, such as the 5′ UTR or coding sequences, to influence gene expression.
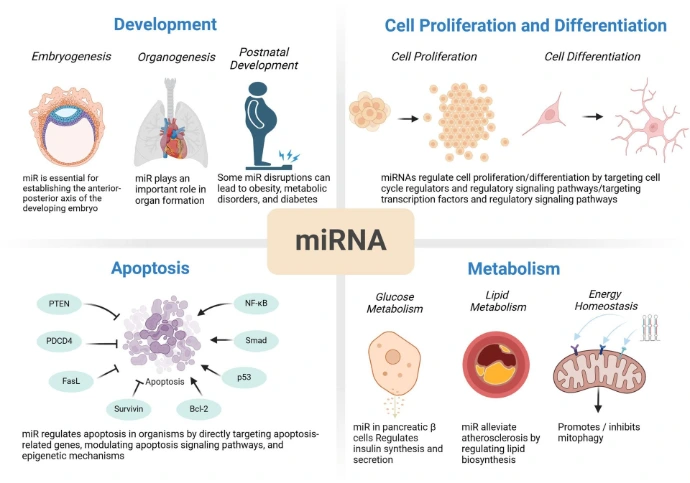
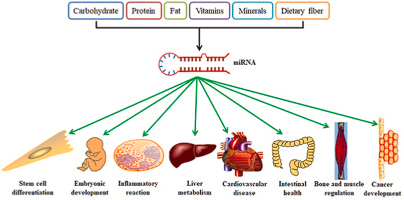
The regulatory effects of miRNAs are context-dependent, varying across cell types, developmental stages, and environmental conditions. They can act as oncogenes or tumor suppressors depending on their targets and the cellular environment. miRNAs play key roles in controlling fundamental biological processes, including development, cell proliferation and differentiation, apoptosis, and metabolism.
Development
Development is a complex and tightly regulated process involving coordinated gene networks and signaling pathways. miRNAs play a crucial role in controlling developmental processes, from embryogenesis to postnatal growth. They regulate development by directly targeting key genes or modulating signaling cascades for example, miR-181 influences T cell development by targeting PTEN, while miR-1 regulates cardiac muscle development by targeting Hand2.
miRNAs also interact with epigenetic mechanisms, affecting the activity of enzymes such as DNA methyltransferases and histone modifiers. For instance, the miR-29 family targets DNMTs, promoting DNA hypomethylation and activating developmental genes. Understanding these interactions is essential for deciphering development and offers potential avenues for therapeutic interventions in developmental disorders.
miRNA Regulation of Organogenesis
miRNAs are critical regulators of organ formation during embryonic development, guiding the differentiation, proliferation, and maturation of cells across multiple organ systems.
Heart: miR-1, miR-133, and miR-208 control cardiomyocyte development and cardiac gene expression. Dysregulation can lead to congenital heart defects or myocardial infarction. miR-1/133a clusters fine-tune myocardin levels to specify cardiomyogenic lineages.
Nervous System: miRNAs such as miR-124, miR-25, and let-7 regulate neural progenitor differentiation, metabolic states, and neuronal maturation. miRNAs like miR-137-3p and miR-153-3p co-target key transcription factors (Neurod1) and genes (Src) to coordinate neurodevelopment.
Skeleton: miR-138, miR-140, and miR-29 influence osteoblast and chondrocyte differentiation, modulating bone formation and skeletal homeostasis.
Lungs: miR-302/367, miR-17-92, miR-200 family, miR-204, and miR-210 regulate progenitor proliferation, branching morphogenesis, epithelial differentiation, and surfactant protein expression.
Kidneys: miRNAs including miR-17 and miR-200 fine-tune nephrogenesis, nephron formation, and tubulogenesis, interacting with Wnt/β-catenin, Notch, and TGF-β signaling pathways. Dysregulation is linked to congenital anomalies and Wilms’ tumor.
Organogenesis is tightly controlled by complex miRNA networks, transcription factors, signaling pathways, and ceRNAs, which together modulate gene expression post-transcriptionally. Understanding these regulatory interactions provides insights into developmental biology and highlights miRNAs as potential therapeutic targets.